Artikelen
Artikelen SNNDV nascholing november 2018 - Next generation melanoma pathology (2019-02)
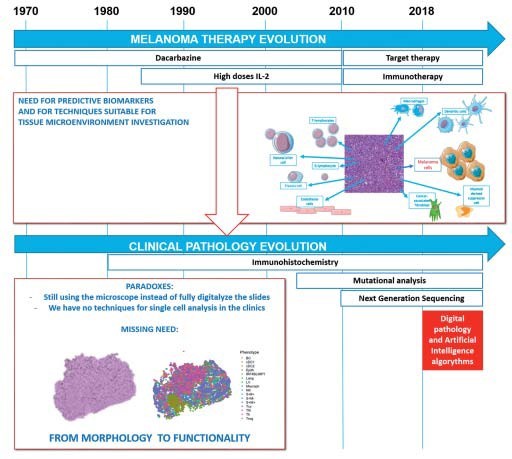
From morphology to functionality
Artikel in PDF
F.M. Bosisio
The fight against cancer has seen a quite dramatic mentality change in the last century. From the search for a mythological holy grail, a powerful drug that could defeat all types of cancer in every patient, we have come nowadays to the realization that a truly effective way to defeat cancer is personalized therapy. This consists in a precise tailoring of the treatment not only according to the type of cancer, but also to the specific characteristics of the patient. Providing the right therapy to each cancer patient heavily relies on an interdisciplinary interaction between surgeons, oncologists and pathologists. While not readily visible to patients, pathological assessments remain essential not only to determine the correct diagnosis, but also to assess the prognosis of the patient and the success of a specific therapeutic plan.
The last ten years have witnessed incredible progresses in melanoma therapy with the introduction of immunotherapy in clinical practice. Nevertheless, there are still a lot of open questions. Only a minority of patients responds to these drugs, and even the ones that respond can develop secondary resistance in 20-30% of the cases. [1-3] Plus, these drugs are burdened by high costs and adverse reactions due to an hyperactive immune response. [4,5] Therefore, there is the need for a deeper understanding of the mechanisms of the immune response in order to find biomarkers that could predict which group of patients will have a real benefit from them.
An intricate topic to investigate
It’s no wonder that these new therapeutic possibilities create new challenges considering that their target, the immune microenvironment of the tumor, is a complex entity. First of all, because it has high variability in space and time. Being a dynamic system, different tumor localizations can have significant differences even within the same patient, and during the course of the disease we could witness alternating waves of activation and exhaustion. In second instance, its complexity is also based on its composition: a huge number of different cell types, each with a specific immunostimulating or immunosuppressive role, that if modified by a drug can influence in turn to modifications of the immune status of all the cells they are respectively interacting with, in an immunological “domino” effect. Moreover, independently of their immune- suppressive/stimulating role, these different cell types can be active, therefore participating to the immune response, or inactive/exhausted, and being unable to have an effect against or in favor of the tumor, even if morphologically present in the histological section. [6] Potential biomarkers that could predict the response to immunotherapy have been identified, such as the mutational load and neoantigen expression, PD-L1 expression, the presence and location of tumor infiltr ting lymphocytes and the activity of the interferon-g signaling pathway in tumor cells. [7] Nevertheless, at the moment, no biomarker is widely accepted for routine clinical use. The insufficient predictive power of these biomarkers could be, again, explained by the partial view that they take over such a complex microenvironment. A vast portion of the studies looking for predictive biomarkers finds predictive parameters in peripheral blood. [8] The main reason is that peripheral blood is easily accessible and that, being a liquid tissue, can be analyzed for multiple parameters at the same time with a fair variety of techniques. Markers found in peripheral blood should reflect the situation at the tumor site. However, there are evidences for substantial differences between parameters in peripheral blood and at the tumor site, such as, for example, the functional status of circulating inflammatory cells and their tissue-based counterparts. [9] It is intuitive that it is the beh vior of the inflammatory cells near the tumor and not that of circulating inflammatory cells that will determine the response to immunotherapy. Finally, for the study of immune cells, it was highly recommended that modifications of the state of the cells in the original tissue should be minimized, in order to mimic the reality and not to favour a specific cell type and losing information about other cell types, and this is better done if the tissue status is blocked through fixation with formalyne. [10] On the other hand, to investigate multiple parameters directly on the tissue of the patient presents multiple technical challenges. A proper investigation of the tumor microenvironment requires a functional, multi-parameter single cell analysis which includes insights in spatial distribution. In few words, we need to get a picture of the immune contexture (type, amount and relative location of the immune cells) and add the functional information to the phenotypical characterization (activation status of the different cell types) if we want to find reliable parameters of response to immunotherapy. In order to achieve this goal, we need more powerful analytic tools. While nextgeneration sequencing procedures to identify genetic aberrations have witnessed an enormous revolution over the last decade, current standard pathological methods are significantly lagging behind to cope with the increasing numbers of putative biomarkers that could lead to more precise diagnostics and better therapy selections. Surprisingly, the techniques applied for general pathology (i.e. immunohistochemistry) have not kept pace: in a world in which a multi-parameter analysis at single cell resolution is becoming the gold standard, pathologists still apply old methods that allow the simultaneous assessment of 1-2 proteins in a single tissue slide, which are subsequently evaluated in a semiquantitative way. [11] Importantly, the number of putative biomarkers identified in research settings keeps growing every day, but integrating such findings in clinical practice remains challenging. [12]
New tools to the rescue!
In the latest years, several methods for single cell-analysis have been implemented in order to obtain a high resolution landscape of the tumor microenvironment, as highlighted in one dedicated issue of Nature in 2017. [13,14] Nevertheless, most of these methods rely on dissociation of the cells from fresh material. This approach makes to study biopsies with very limited material, that must be fully dedicated to the pathological diagnosis, most of the times impossible. Moreover, high resolution means also to be able to characterize not only the heterogeneity of the immune infiltrate but also to define the spatial distribution of each component within it. [15] Spatial distribution adds information that allows to make inferences about the interactions of the cells in the tissue, a feature that is completely lost with tissue dissociation. Strikingly, most of these methods are either incredibly expensive, require complicated procedures or have many technical and practical disadvantages, severely limiting the analysis of large cohorts of tumor samples. Some of these disadvantages are, for example, the loss of cells between the cycles for bleaching/quen- ching techniques; the steric hindrance between sequentially deposited antibodies that target nearby epitopes and that may mask the detection of the epitopes in later staining cycles when tyramide-based amplification methods are used; the possibility to investigate only regions of interest and not the whole slide due to the long scanning times; the possibility to work only on frozen material, not always available for every patient, for some of these methods. [16,17] Finally, these techniques imply destruction of the tissue section during the analysis, making it not available for further investigations. Therefore, until recently, an effective method to stain tissue sections with multiple markers at the single cell resolution was lacking.
First steps towards a revolution
In the last years, a collaboration between the KU Leuven and the University of Milan-Bicocca has co-developed a novel tool, called “Multiple Iteractive Labeling by Antibody Neodeposition” (M.I.L.A.N.), to finally deal with this issue. [18,19] Using these methods we were able to obtain a high resolution picture that we used to obtain preliminary data to untangle the microenvironment in the “brisk” and “non brisk” morphologic categories of tumor infiltrating lymphocytes, not only quantitatively but also functionally through neighbor analysis. Not only we obtained a functional classification of the T cell infiltrates that could be considered more accurate than what was previously done in literature, but we also explored which other immune cells were verisimilarly interacting with active and exhausted T cells. Analyzing 40 immune markers we were able to deeply phenotype the infiltrate (not only the main T cell subsets, but also B cells, macrophages and different types of dendritic cells) in active and exhausted cases and detect the most frequent interactions and avoidance relationships between them.
Conclusion
In conclusion, a thorough evaluation of the tumor microenvironment done directly on tissue section from the patient through an high-dimensional multiplex technique is needed in order to depict the complexity of the immune microenvironment and reach a satisfactory predictive power in clinical practice. Similar tools on peripheral blood or frozen tissue are at the moment merely centered on predicting the response to immunotherapy, but should evolve in the future to represent a tool to decide which type of treatment the patient should receive. By applying it not only at the beginning of the therapeutic path of the patient but also during the course of the treatment, it will represent a tool to follow the evolving tumoral landscape and it will allow to monitor response of the patient to the therapeutic protocols, thereby fulfilling the need for a completely personalized therapy for the patient.
Literatuur
1. Ribas A, Flaherty KT. Gauging the long-term benefits of ipilimumab in melanoma. J Clin Oncol 2015;33(17):1865-6.
2. Ribas A, Hamid O, Daud A, et al. association of pembrolizumab with tumor response and survival among patients with advanced mela- noma. JAMA 2016;315(15):1600-9.
3. Robert C, Long GV, Brady B, et al. Nivolumab in previously untreated melanoma without BRAF mutation. N Engl J Med 2015;372(4):320-30. Wang DY, Salem JE, Cohen JV, et al. Fatal toxic effects associated with immune checkpoint inhibitors: A Systematic Review and Meta-analy- sis. JAMA Oncol 2018 [Epub ahead of print]
4. Postow MA, Hellmann MD. Adverse Events Associated with Immune Checkpoint Blockade. N Engl J Med. 2018 Mar 22;378(12):1165.
5. Bosisio FM, van den Oord JJ. Immunoplasticity in cutaneous mela- noma: beyond pure morphology. Virchows Arch. 2017;470(4):357-369.
6. Schadendorf D, van Akkooi ACJ, Berking C, et al. Melanoma. The Lancet 2018;392(10151): 971.
7. Ascierto PA, Agarwala SS, Ciliberto G, et al. Future perspectives in melanoma research “Melanoma Bridge”, Napoli, November 30th-3rd December 2016. J Transl Med 2017;15(1):236.
8. Buggert M, Nguyen S, Salgado-Montes de Oca G, et al. Identification and characterization of HIV-specific resident memory CD8+ T cells in human lymphoid tissue. Sci Immunol 2018;3(24).
9. Giladi A, Amit I. Immunology, one cell at a time. Nature 2017;547(7661):27-9.
10. Caie PD, Harrison DJ. Next-generation pathology. Methods Mol Biol 2018;1386:61-72.
11. Henry NL, Hayes DF. Cancer biomarkers. Mol Oncol 2012;6:140-6. Single-cell biology. Nature 2017;547:19.
De complete literatuurlijst is, vanaf drie weken na publicatie in dit tijdschrift, te vinden op www.nvdv.nl.
Samenvatting
De snelle evolutie in de behandeling van melanomen dwingt de pathologie een nieuw tijdperk in te gaan waarin de evaluatie van meerdere biomarkers op individueel celniveau gekoppeld wordt met de nieuwe mogelijkheden van artificiële intelligentie. Kan pathologie de kliniek volgen?
Summary
The rapidly evolving landscape of melanoma therapy is urging pathology to enter in a new era, in which the evaluation of multiple biomarkers at single cell level is coupled with the new possibilities that artificial intelligence can offer. Is pathology ready to follow the clinic?
Trefwoorden
melanoom – pathologie – beeldanalyse – multiplex – immunotherapie
Keywords
melanoma – pathology – image analysis – multiplex – immunotherapy
Correspondentieadres
Francesca Bosisio
E-mail: francesca.bosisio@uzleuven.be